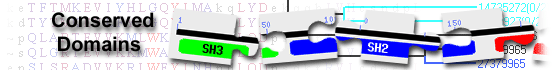leucine zipper protein 2 isoform 3 [Homo sapiens]
coiled-coil domain-containing protein( domain architecture ID 1000037)
coiled-coil domain-containing protein contains a region with alpha-helical coiled-coil sequence signatures that is being annotated by a variety of protein family models, not necessarily indicating family membership
List of domain hits

| Name | Accession | Description | Interval | E-value | ||||
| EnvC super family | cl34844 | Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, ... |
21-169 | 1.15e-06 | ||||
Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, cell division, chromosome partitioning]; The actual alignment was detected with superfamily member COG4942: Pssm-ID: 443969 [Multi-domain] Cd Length: 377 Bit Score: 49.38 E-value: 1.15e-06
|
||||||||
| Name | Accession | Description | Interval | E-value | ||||
| EnvC | COG4942 | Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, ... |
21-169 | 1.15e-06 | ||||
Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 443969 [Multi-domain] Cd Length: 377 Bit Score: 49.38 E-value: 1.15e-06
|
||||||||
| SMC_prok_B | TIGR02168 | chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of ... |
25-167 | 2.21e-06 | ||||
chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of chromosomes) proteins bind DNA and act in organizing and segregating chromosomes for partition. SMC proteins are found in bacteria, archaea, and eukaryotes. This family represents the SMC protein of most bacteria. The smc gene is often associated with scpB (TIGR00281) and scpA genes, where scp stands for segregation and condensation protein. SMC was shown (in Caulobacter crescentus) to be induced early in S phase but present and bound to DNA throughout the cell cycle. [Cellular processes, Cell division, DNA metabolism, Chromosome-associated proteins] Pssm-ID: 274008 [Multi-domain] Cd Length: 1179 Bit Score: 49.28 E-value: 2.21e-06
|
||||||||
| PRK00409 | PRK00409 | recombination and DNA strand exchange inhibitor protein; Reviewed |
60-166 | 1.64e-04 | ||||
recombination and DNA strand exchange inhibitor protein; Reviewed Pssm-ID: 234750 [Multi-domain] Cd Length: 782 Bit Score: 43.28 E-value: 1.64e-04
|
||||||||
| ClyA_Cry6Aa-like | cd22656 | Bacillus thuringiensis crystal 6Aa (Cry6Aa) toxin, and similar proteins; This model includes ... |
25-152 | 2.10e-04 | ||||
Bacillus thuringiensis crystal 6Aa (Cry6Aa) toxin, and similar proteins; This model includes pesticidal Cry6Aa toxin from Bacillus thuringiensis, one of the many parasporal crystal (Cry) toxins produced during the sporulation phase of growth. Many of these proteins are toxic to numerous insect species and have been effectively used as proteinaceous insecticides to directly kill insect pests; some have been used to control insect growth on transgenic agricultural plants. Cry6Aa exists as a protoxin, which is activated by cleavage using trypsin. Structure studies for Cry6Aa support a mechanism of action by pore formation, similar to cytolysin A (ClyA)-type alpha pore-forming toxins (alpha-PFTs) such as HblB, and bioassay and mutation studies show that Cry6Aa is an active pore-forming toxin. Cry6Aa shows atypical features compared to other members of alpha-PFTs, including internal repeat sequences and small loop regions within major alpha helices. Pssm-ID: 439154 [Multi-domain] Cd Length: 309 Bit Score: 42.36 E-value: 2.10e-04
|
||||||||
| FapA | pfam03961 | Flagellar Assembly Protein A beta solenoid domain; This entry represents the C-terminal beta ... |
77-152 | 1.26e-03 | ||||
Flagellar Assembly Protein A beta solenoid domain; This entry represents the C-terminal beta solenoid domain of FapA and its homologs. Members of this family include FapA (flagellar assembly protein A) found in Vibrio vulnificus. The synthesis of flagella allows bacteria to respond to chemotaxis by facilitating motility. Studies examining the role of FapA show that the loss or delocalization of FapA results in a complete failure of the flagellar biosynthesis and motility in response to glucose mediated chemotaxis. The polar localization of FapA is required for flagellar synthesis, and dephosphorylated EIIAGlc (Glucose-permease IIA component) inhibited the polar localization of FapA through direct interaction. This entry shows similarity to pfam03775 suggesting a similar functional role. Pssm-ID: 461111 [Multi-domain] Cd Length: 272 Bit Score: 39.59 E-value: 1.26e-03
|
||||||||
| Name | Accession | Description | Interval | E-value | ||||
| EnvC | COG4942 | Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, ... |
21-169 | 1.15e-06 | ||||
Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 443969 [Multi-domain] Cd Length: 377 Bit Score: 49.38 E-value: 1.15e-06
|
||||||||
| SMC_prok_B | TIGR02168 | chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of ... |
25-167 | 2.21e-06 | ||||
chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of chromosomes) proteins bind DNA and act in organizing and segregating chromosomes for partition. SMC proteins are found in bacteria, archaea, and eukaryotes. This family represents the SMC protein of most bacteria. The smc gene is often associated with scpB (TIGR00281) and scpA genes, where scp stands for segregation and condensation protein. SMC was shown (in Caulobacter crescentus) to be induced early in S phase but present and bound to DNA throughout the cell cycle. [Cellular processes, Cell division, DNA metabolism, Chromosome-associated proteins] Pssm-ID: 274008 [Multi-domain] Cd Length: 1179 Bit Score: 49.28 E-value: 2.21e-06
|
||||||||
| Smc | COG1196 | Chromosome segregation ATPase Smc [Cell cycle control, cell division, chromosome partitioning]; ... |
25-166 | 1.08e-05 | ||||
Chromosome segregation ATPase Smc [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 440809 [Multi-domain] Cd Length: 983 Bit Score: 46.85 E-value: 1.08e-05
|
||||||||
| SMC_prok_B | TIGR02168 | chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of ... |
22-166 | 1.47e-05 | ||||
chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of chromosomes) proteins bind DNA and act in organizing and segregating chromosomes for partition. SMC proteins are found in bacteria, archaea, and eukaryotes. This family represents the SMC protein of most bacteria. The smc gene is often associated with scpB (TIGR00281) and scpA genes, where scp stands for segregation and condensation protein. SMC was shown (in Caulobacter crescentus) to be induced early in S phase but present and bound to DNA throughout the cell cycle. [Cellular processes, Cell division, DNA metabolism, Chromosome-associated proteins] Pssm-ID: 274008 [Multi-domain] Cd Length: 1179 Bit Score: 46.59 E-value: 1.47e-05
|
||||||||
| CwlO1 | COG3883 | Uncharacterized N-terminal coiled-coil domain of peptidoglycan hydrolase CwlO [Function ... |
19-168 | 1.49e-05 | ||||
Uncharacterized N-terminal coiled-coil domain of peptidoglycan hydrolase CwlO [Function unknown]; Pssm-ID: 443091 [Multi-domain] Cd Length: 379 Bit Score: 45.98 E-value: 1.49e-05
|
||||||||
| Smc | COG1196 | Chromosome segregation ATPase Smc [Cell cycle control, cell division, chromosome partitioning]; ... |
21-169 | 1.67e-05 | ||||
Chromosome segregation ATPase Smc [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 440809 [Multi-domain] Cd Length: 983 Bit Score: 46.47 E-value: 1.67e-05
|
||||||||
| Smc | COG1196 | Chromosome segregation ATPase Smc [Cell cycle control, cell division, chromosome partitioning]; ... |
24-169 | 2.44e-05 | ||||
Chromosome segregation ATPase Smc [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 440809 [Multi-domain] Cd Length: 983 Bit Score: 45.70 E-value: 2.44e-05
|
||||||||
| GumC | COG3206 | Exopolysaccharide export protein/domain GumC/Wzc1 [Cell wall/membrane/envelope biogenesis]; |
21-196 | 5.84e-05 | ||||
Exopolysaccharide export protein/domain GumC/Wzc1 [Cell wall/membrane/envelope biogenesis]; Pssm-ID: 442439 [Multi-domain] Cd Length: 687 Bit Score: 44.62 E-value: 5.84e-05
|
||||||||
| EnvC | COG4942 | Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, ... |
19-166 | 1.28e-04 | ||||
Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 443969 [Multi-domain] Cd Length: 377 Bit Score: 43.21 E-value: 1.28e-04
|
||||||||
| YhaN | COG4717 | Uncharacterized conserved protein YhaN, contains AAA domain [Function unknown]; |
19-159 | 1.39e-04 | ||||
Uncharacterized conserved protein YhaN, contains AAA domain [Function unknown]; Pssm-ID: 443752 [Multi-domain] Cd Length: 641 Bit Score: 43.22 E-value: 1.39e-04
|
||||||||
| PRK00409 | PRK00409 | recombination and DNA strand exchange inhibitor protein; Reviewed |
60-166 | 1.64e-04 | ||||
recombination and DNA strand exchange inhibitor protein; Reviewed Pssm-ID: 234750 [Multi-domain] Cd Length: 782 Bit Score: 43.28 E-value: 1.64e-04
|
||||||||
| COG4913 | COG4913 | Uncharacterized conserved protein, contains a C-terminal ATPase domain [Function unknown]; |
22-166 | 1.67e-04 | ||||
Uncharacterized conserved protein, contains a C-terminal ATPase domain [Function unknown]; Pssm-ID: 443941 [Multi-domain] Cd Length: 1089 Bit Score: 43.37 E-value: 1.67e-04
|
||||||||
| ClyA_Cry6Aa-like | cd22656 | Bacillus thuringiensis crystal 6Aa (Cry6Aa) toxin, and similar proteins; This model includes ... |
25-152 | 2.10e-04 | ||||
Bacillus thuringiensis crystal 6Aa (Cry6Aa) toxin, and similar proteins; This model includes pesticidal Cry6Aa toxin from Bacillus thuringiensis, one of the many parasporal crystal (Cry) toxins produced during the sporulation phase of growth. Many of these proteins are toxic to numerous insect species and have been effectively used as proteinaceous insecticides to directly kill insect pests; some have been used to control insect growth on transgenic agricultural plants. Cry6Aa exists as a protoxin, which is activated by cleavage using trypsin. Structure studies for Cry6Aa support a mechanism of action by pore formation, similar to cytolysin A (ClyA)-type alpha pore-forming toxins (alpha-PFTs) such as HblB, and bioassay and mutation studies show that Cry6Aa is an active pore-forming toxin. Cry6Aa shows atypical features compared to other members of alpha-PFTs, including internal repeat sequences and small loop regions within major alpha helices. Pssm-ID: 439154 [Multi-domain] Cd Length: 309 Bit Score: 42.36 E-value: 2.10e-04
|
||||||||
| SMC_prok_B | TIGR02168 | chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of ... |
20-163 | 2.44e-04 | ||||
chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of chromosomes) proteins bind DNA and act in organizing and segregating chromosomes for partition. SMC proteins are found in bacteria, archaea, and eukaryotes. This family represents the SMC protein of most bacteria. The smc gene is often associated with scpB (TIGR00281) and scpA genes, where scp stands for segregation and condensation protein. SMC was shown (in Caulobacter crescentus) to be induced early in S phase but present and bound to DNA throughout the cell cycle. [Cellular processes, Cell division, DNA metabolism, Chromosome-associated proteins] Pssm-ID: 274008 [Multi-domain] Cd Length: 1179 Bit Score: 42.74 E-value: 2.44e-04
|
||||||||
| YhaN | COG4717 | Uncharacterized conserved protein YhaN, contains AAA domain [Function unknown]; |
22-166 | 2.52e-04 | ||||
Uncharacterized conserved protein YhaN, contains AAA domain [Function unknown]; Pssm-ID: 443752 [Multi-domain] Cd Length: 641 Bit Score: 42.45 E-value: 2.52e-04
|
||||||||
| DR0291 | COG1579 | Predicted nucleic acid-binding protein DR0291, contains C4-type Zn-ribbon domain [General ... |
26-166 | 3.54e-04 | ||||
Predicted nucleic acid-binding protein DR0291, contains C4-type Zn-ribbon domain [General function prediction only]; Pssm-ID: 441187 [Multi-domain] Cd Length: 236 Bit Score: 41.06 E-value: 3.54e-04
|
||||||||
| PRK05431 | PRK05431 | seryl-tRNA synthetase; Provisional |
72-155 | 3.65e-04 | ||||
seryl-tRNA synthetase; Provisional Pssm-ID: 235461 [Multi-domain] Cd Length: 425 Bit Score: 41.59 E-value: 3.65e-04
|
||||||||
| COG4372 | COG4372 | Uncharacterized protein, contains DUF3084 domain [Function unknown]; |
39-169 | 4.48e-04 | ||||
Uncharacterized protein, contains DUF3084 domain [Function unknown]; Pssm-ID: 443500 [Multi-domain] Cd Length: 370 Bit Score: 41.43 E-value: 4.48e-04
|
||||||||
| PRK11637 | PRK11637 | AmiB activator; Provisional |
19-162 | 4.96e-04 | ||||
AmiB activator; Provisional Pssm-ID: 236942 [Multi-domain] Cd Length: 428 Bit Score: 41.22 E-value: 4.96e-04
|
||||||||
| SMC_prok_B | TIGR02168 | chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of ... |
25-170 | 6.59e-04 | ||||
chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of chromosomes) proteins bind DNA and act in organizing and segregating chromosomes for partition. SMC proteins are found in bacteria, archaea, and eukaryotes. This family represents the SMC protein of most bacteria. The smc gene is often associated with scpB (TIGR00281) and scpA genes, where scp stands for segregation and condensation protein. SMC was shown (in Caulobacter crescentus) to be induced early in S phase but present and bound to DNA throughout the cell cycle. [Cellular processes, Cell division, DNA metabolism, Chromosome-associated proteins] Pssm-ID: 274008 [Multi-domain] Cd Length: 1179 Bit Score: 41.20 E-value: 6.59e-04
|
||||||||
| HEC1 | COG5185 | Chromosome segregation protein NDC80, interacts with SMC proteins [Cell cycle control, cell ... |
22-169 | 6.89e-04 | ||||
Chromosome segregation protein NDC80, interacts with SMC proteins [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 444066 [Multi-domain] Cd Length: 594 Bit Score: 41.10 E-value: 6.89e-04
|
||||||||
| SMC_prok_A | TIGR02169 | chromosome segregation protein SMC, primarily archaeal type; SMC (structural maintenance of ... |
21-187 | 9.52e-04 | ||||
chromosome segregation protein SMC, primarily archaeal type; SMC (structural maintenance of chromosomes) proteins bind DNA and act in organizing and segregating chromosomes for partition. SMC proteins are found in bacteria, archaea, and eukaryotes. It is found in a single copy and is homodimeric in prokaryotes, but six paralogs (excluded from this family) are found in eukarotes, where SMC proteins are heterodimeric. This family represents the SMC protein of archaea and a few bacteria (Aquifex, Synechocystis, etc); the SMC of other bacteria is described by TIGR02168. The N- and C-terminal domains of this protein are well conserved, but the central hinge region is skewed in composition and highly divergent. [Cellular processes, Cell division, DNA metabolism, Chromosome-associated proteins] Pssm-ID: 274009 [Multi-domain] Cd Length: 1164 Bit Score: 40.82 E-value: 9.52e-04
|
||||||||
| YhaN | COG4717 | Uncharacterized conserved protein YhaN, contains AAA domain [Function unknown]; |
25-169 | 9.92e-04 | ||||
Uncharacterized conserved protein YhaN, contains AAA domain [Function unknown]; Pssm-ID: 443752 [Multi-domain] Cd Length: 641 Bit Score: 40.52 E-value: 9.92e-04
|
||||||||
| PRK03918 | PRK03918 | DNA double-strand break repair ATPase Rad50; |
24-171 | 1.21e-03 | ||||
DNA double-strand break repair ATPase Rad50; Pssm-ID: 235175 [Multi-domain] Cd Length: 880 Bit Score: 40.43 E-value: 1.21e-03
|
||||||||
| FapA | pfam03961 | Flagellar Assembly Protein A beta solenoid domain; This entry represents the C-terminal beta ... |
77-152 | 1.26e-03 | ||||
Flagellar Assembly Protein A beta solenoid domain; This entry represents the C-terminal beta solenoid domain of FapA and its homologs. Members of this family include FapA (flagellar assembly protein A) found in Vibrio vulnificus. The synthesis of flagella allows bacteria to respond to chemotaxis by facilitating motility. Studies examining the role of FapA show that the loss or delocalization of FapA results in a complete failure of the flagellar biosynthesis and motility in response to glucose mediated chemotaxis. The polar localization of FapA is required for flagellar synthesis, and dephosphorylated EIIAGlc (Glucose-permease IIA component) inhibited the polar localization of FapA through direct interaction. This entry shows similarity to pfam03775 suggesting a similar functional role. Pssm-ID: 461111 [Multi-domain] Cd Length: 272 Bit Score: 39.59 E-value: 1.26e-03
|
||||||||
| SMC_prok_A | TIGR02169 | chromosome segregation protein SMC, primarily archaeal type; SMC (structural maintenance of ... |
22-166 | 1.32e-03 | ||||
chromosome segregation protein SMC, primarily archaeal type; SMC (structural maintenance of chromosomes) proteins bind DNA and act in organizing and segregating chromosomes for partition. SMC proteins are found in bacteria, archaea, and eukaryotes. It is found in a single copy and is homodimeric in prokaryotes, but six paralogs (excluded from this family) are found in eukarotes, where SMC proteins are heterodimeric. This family represents the SMC protein of archaea and a few bacteria (Aquifex, Synechocystis, etc); the SMC of other bacteria is described by TIGR02168. The N- and C-terminal domains of this protein are well conserved, but the central hinge region is skewed in composition and highly divergent. [Cellular processes, Cell division, DNA metabolism, Chromosome-associated proteins] Pssm-ID: 274009 [Multi-domain] Cd Length: 1164 Bit Score: 40.44 E-value: 1.32e-03
|
||||||||
| SMC_prok_B | TIGR02168 | chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of ... |
21-158 | 1.99e-03 | ||||
chromosome segregation protein SMC, common bacterial type; SMC (structural maintenance of chromosomes) proteins bind DNA and act in organizing and segregating chromosomes for partition. SMC proteins are found in bacteria, archaea, and eukaryotes. This family represents the SMC protein of most bacteria. The smc gene is often associated with scpB (TIGR00281) and scpA genes, where scp stands for segregation and condensation protein. SMC was shown (in Caulobacter crescentus) to be induced early in S phase but present and bound to DNA throughout the cell cycle. [Cellular processes, Cell division, DNA metabolism, Chromosome-associated proteins] Pssm-ID: 274008 [Multi-domain] Cd Length: 1179 Bit Score: 39.65 E-value: 1.99e-03
|
||||||||
| PRK03918 | PRK03918 | DNA double-strand break repair ATPase Rad50; |
25-169 | 2.18e-03 | ||||
DNA double-strand break repair ATPase Rad50; Pssm-ID: 235175 [Multi-domain] Cd Length: 880 Bit Score: 39.66 E-value: 2.18e-03
|
||||||||
| COG4913 | COG4913 | Uncharacterized conserved protein, contains a C-terminal ATPase domain [Function unknown]; |
19-153 | 2.64e-03 | ||||
Uncharacterized conserved protein, contains a C-terminal ATPase domain [Function unknown]; Pssm-ID: 443941 [Multi-domain] Cd Length: 1089 Bit Score: 39.51 E-value: 2.64e-03
|
||||||||
| EnvC | COG4942 | Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, ... |
81-169 | 3.03e-03 | ||||
Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 443969 [Multi-domain] Cd Length: 377 Bit Score: 38.98 E-value: 3.03e-03
|
||||||||
| COG4913 | COG4913 | Uncharacterized conserved protein, contains a C-terminal ATPase domain [Function unknown]; |
21-151 | 3.31e-03 | ||||
Uncharacterized conserved protein, contains a C-terminal ATPase domain [Function unknown]; Pssm-ID: 443941 [Multi-domain] Cd Length: 1089 Bit Score: 39.13 E-value: 3.31e-03
|
||||||||
| DUF3584 | pfam12128 | Protein of unknown function (DUF3584); This protein is found in bacteria and eukaryotes. ... |
19-166 | 5.31e-03 | ||||
Protein of unknown function (DUF3584); This protein is found in bacteria and eukaryotes. Proteins in this family are typically between 943 to 1234 amino acids in length. This family contains a P-loop motif suggesting it is a nucleotide binding protein. It may be involved in replication. Pssm-ID: 432349 [Multi-domain] Cd Length: 1191 Bit Score: 38.67 E-value: 5.31e-03
|
||||||||
| CwlO1 | COG3883 | Uncharacterized N-terminal coiled-coil domain of peptidoglycan hydrolase CwlO [Function ... |
77-154 | 6.23e-03 | ||||
Uncharacterized N-terminal coiled-coil domain of peptidoglycan hydrolase CwlO [Function unknown]; Pssm-ID: 443091 [Multi-domain] Cd Length: 379 Bit Score: 37.89 E-value: 6.23e-03
|
||||||||
| Smc | COG1196 | Chromosome segregation ATPase Smc [Cell cycle control, cell division, chromosome partitioning]; ... |
21-169 | 7.46e-03 | ||||
Chromosome segregation ATPase Smc [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 440809 [Multi-domain] Cd Length: 983 Bit Score: 37.99 E-value: 7.46e-03
|
||||||||
| ATG17_like | pfam04108 | Autophagy protein ATG17-like domain; This domain is found in the autophagy-related proteins ... |
19-167 | 7.56e-03 | ||||
Autophagy protein ATG17-like domain; This domain is found in the autophagy-related proteins ATG17 and ATG11, conserved across eukaryotes. ATG17 forms a complex with ATG29 and ATG31, critical for both PAS (preautophagosomal structure) formation and autophagy. Together with ATG13, it is required for ATG1 kinase activation. ATG11 is a scaffold protein required for the cytoplasm-to-vacuole targeting (Cvt) pathway during starvation and to recruit ATG proteins to the pre-autophagosome. It is also required for ATG1 kinase activation. In many eukaryotes, ATG11 (the orthologue in mammals is RB1-inducible coiled-coil protein 1 (RB1CC1) and in S. pombe is Taz1-interacting factor 1 (taf1)) is essential for bulk autophagy, except in S.cerevisiae. ATG17 and ATG11 are large similar proteins, both predicted to be almost entirely helical, containing conserved coiled-coil regions and lack obvious functional motifs. Pssm-ID: 427715 [Multi-domain] Cd Length: 360 Bit Score: 37.75 E-value: 7.56e-03
|
||||||||
| Mplasa_alph_rch | TIGR04523 | helix-rich Mycoplasma protein; Members of this family occur strictly within a subset of ... |
22-169 | 7.94e-03 | ||||
helix-rich Mycoplasma protein; Members of this family occur strictly within a subset of Mycoplasma species. Members average 750 amino acids in length, including signal peptide. Sequences are predicted (Jpred 3) to be almost entirely alpha-helical. These sequences show strong periodicity (consistent with long alpha helical structures) and low complexity rich in D,E,N,Q, and K. Genes encoding these proteins are often found in tandem. The function is unknown. Pssm-ID: 275316 [Multi-domain] Cd Length: 745 Bit Score: 37.69 E-value: 7.94e-03
|
||||||||
| EnvC | COG4942 | Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, ... |
60-152 | 9.54e-03 | ||||
Septal ring factor EnvC, activator of murein hydrolases AmiA and AmiB [Cell cycle control, cell division, chromosome partitioning]; Pssm-ID: 443969 [Multi-domain] Cd Length: 377 Bit Score: 37.44 E-value: 9.54e-03
|
||||||||
| Blast search parameters | ||||
|