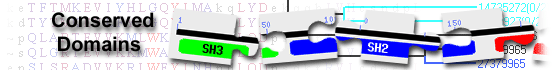ubiquitin carboxyl-terminal hydrolase 19 isoform X2 [Homo sapiens]
ubiquitin specific protease( domain architecture ID 12950362)
ubiquitin specific protease (such as USP19) regulates cell cycle progression and is involved in the cellular response to different types of stress, including the unfolded protein response (UPR), hypoxia and muscle atrophy.
List of domain hits

| Name | Accession | Description | Interval | E-value | |||||||||||
| UBP12 super family | cl35019 | Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; |
597-1319 | 1.21e-118 | |||||||||||
Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; The actual alignment was detected with superfamily member COG5560: Pssm-ID: 227847 [Multi-domain] Cd Length: 823 Bit Score: 391.55 E-value: 1.21e-118
|
|||||||||||||||
| USP19_linker | pfam16602 | Linker region of USP19 deubiquitinase; This region is generally located between a CS domain ... |
477-599 | 5.74e-58 | |||||||||||
Linker region of USP19 deubiquitinase; This region is generally located between a CS domain pfam04969 and the enzymatic UCH domain pfam00582 of USP19 deubiquitinases. This region is predicted to be natively unstructured. Its precise functional role is not known. : Pssm-ID: 465191 Cd Length: 121 Bit Score: 195.52 E-value: 5.74e-58
|
|||||||||||||||
| p23_CS_SGT1_like | cd06466 | p23_like domain similar to the C-terminal CHORD-SGT1 (CS) domain of Sgt1 (suppressor of G2 ... |
390-486 | 4.69e-21 | |||||||||||
p23_like domain similar to the C-terminal CHORD-SGT1 (CS) domain of Sgt1 (suppressor of G2 allele of Skp1). Sgt1 interacts with multiple protein complexes and has the features of a cochaperone. Human (h) Sgt1 interacts with both Hsp70 and Hsp90, and has been shown to bind Hsp90 through its CS domain. Saccharomyces cerevisiae (Sc) Sgt1 is a subunit of both core kinetochore and SCF (Skp1-Cul1-F-box) ubiquitin ligase complexes. Sgt1 is required for pathogen resistance in plants. ScSgt1 is needed for the G1/S and G2/M cell-cycle transitions, and for assembly of the core kinetochore complex (CBF3) via activation of Ctf13, the F-box protein. Binding of Hsp82 (a yeast Hsp90 homologue) to ScSgt1, promotes the binding of Sgt1 to Skp1 and of Skp1 to Ctf13. Some proteins in this group have an SGT1-specific (SGS) domain at the extreme C-terminus. The ScSgt1-SGS domain binds adenylate cyclase. The hSgt1-SGS domain interacts with some S100 family proteins, and studies suggest that the interaction of hSgt1 with Hsp90 and Hsp70 may be regulated by S100A6 in a Ca2+ dependent fashion. This group also includes the p23_like domains of Melusin and NAD(P)H cytochrome b5 (NCB5) oxidoreductase (OR). Melusin is a vertebrate protein which interacts with two splice variants of beta1 integrin, and NCB5OR plays a part in maintaining viable pancreatic beta cells. : Pssm-ID: 107223 Cd Length: 84 Bit Score: 88.80 E-value: 4.69e-21
|
|||||||||||||||
| p23_like | cd06463 | Proteins containing this p23_like domain include p23 and its Saccharomyces cerevisiae (Sc) ... |
120-202 | 2.85e-06 | |||||||||||
Proteins containing this p23_like domain include p23 and its Saccharomyces cerevisiae (Sc) homolog Sba1. Both are co-chaperones for the heat shock protein (Hsp) 90. p23 binds Hsp90 and participates in the folding of a number of Hsp90 clients, including the progesterone receptor. p23 also has a passive chaperoning activity and in addition may participate in prostaglandin synthesis. Both p23 and Sba1p can regulate telomerase activity. This group includes domains similar to the C-terminal CHORD-SGT1 (CS) domain of suppressor of G2 allele of Skp1 (Sgt1). Sgt1 interacts with multiple protein complexes and has the features of a co-chaperone. Human (h) Sgt1 interacts with both Hsp70 and Hsp90, and has been shown to bind Hsp90 through its CS domain. Saccharomyces cerevisiae (Sc) Sgt1 is a subunit of both core kinetochore and SCF (Skp1-Cul1-F-box) ubiquitin ligase complexes. Sgt1 is required for pathogen resistance in plants. This group also includes the p23_like domains of human butyrate-induced transcript 1 (hB-ind1), NUD (nuclear distribution) C, Melusin, and NAD(P)H cytochrome b5 (NCB5) oxidoreductase (OR). hB-ind1 plays a role in the signaling pathway mediated by the small GTPase Rac1, NUDC is needed for nuclear movement, Melusin interacts with two splice variants of beta1 integrin, and NCB5OR plays a part in maintaining viable pancreatic beta cells. : Pssm-ID: 107220 Cd Length: 84 Bit Score: 46.51 E-value: 2.85e-06
|
|||||||||||||||
| Name | Accession | Description | Interval | E-value | |||||||||||
| UBP12 | COG5560 | Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; |
597-1319 | 1.21e-118 | |||||||||||
Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; Pssm-ID: 227847 [Multi-domain] Cd Length: 823 Bit Score: 391.55 E-value: 1.21e-118
|
|||||||||||||||
| USP19_linker | pfam16602 | Linker region of USP19 deubiquitinase; This region is generally located between a CS domain ... |
477-599 | 5.74e-58 | |||||||||||
Linker region of USP19 deubiquitinase; This region is generally located between a CS domain pfam04969 and the enzymatic UCH domain pfam00582 of USP19 deubiquitinases. This region is predicted to be natively unstructured. Its precise functional role is not known. Pssm-ID: 465191 Cd Length: 121 Bit Score: 195.52 E-value: 5.74e-58
|
|||||||||||||||
| Peptidase_C19R | cd02674 | A subfamily of peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1159-1316 | 6.15e-56 | |||||||||||
A subfamily of peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239139 [Multi-domain] Cd Length: 230 Bit Score: 194.04 E-value: 6.15e-56
|
|||||||||||||||
| UCH | pfam00443 | Ubiquitin carboxyl-terminal hydrolase; |
601-788 | 3.19e-48 | |||||||||||
Ubiquitin carboxyl-terminal hydrolase; Pssm-ID: 425685 [Multi-domain] Cd Length: 310 Bit Score: 174.94 E-value: 3.19e-48
|
|||||||||||||||
| p23_CS_SGT1_like | cd06466 | p23_like domain similar to the C-terminal CHORD-SGT1 (CS) domain of Sgt1 (suppressor of G2 ... |
390-486 | 4.69e-21 | |||||||||||
p23_like domain similar to the C-terminal CHORD-SGT1 (CS) domain of Sgt1 (suppressor of G2 allele of Skp1). Sgt1 interacts with multiple protein complexes and has the features of a cochaperone. Human (h) Sgt1 interacts with both Hsp70 and Hsp90, and has been shown to bind Hsp90 through its CS domain. Saccharomyces cerevisiae (Sc) Sgt1 is a subunit of both core kinetochore and SCF (Skp1-Cul1-F-box) ubiquitin ligase complexes. Sgt1 is required for pathogen resistance in plants. ScSgt1 is needed for the G1/S and G2/M cell-cycle transitions, and for assembly of the core kinetochore complex (CBF3) via activation of Ctf13, the F-box protein. Binding of Hsp82 (a yeast Hsp90 homologue) to ScSgt1, promotes the binding of Sgt1 to Skp1 and of Skp1 to Ctf13. Some proteins in this group have an SGT1-specific (SGS) domain at the extreme C-terminus. The ScSgt1-SGS domain binds adenylate cyclase. The hSgt1-SGS domain interacts with some S100 family proteins, and studies suggest that the interaction of hSgt1 with Hsp90 and Hsp70 may be regulated by S100A6 in a Ca2+ dependent fashion. This group also includes the p23_like domains of Melusin and NAD(P)H cytochrome b5 (NCB5) oxidoreductase (OR). Melusin is a vertebrate protein which interacts with two splice variants of beta1 integrin, and NCB5OR plays a part in maintaining viable pancreatic beta cells. Pssm-ID: 107223 Cd Length: 84 Bit Score: 88.80 E-value: 4.69e-21
|
|||||||||||||||
| p23_like | cd06463 | Proteins containing this p23_like domain include p23 and its Saccharomyces cerevisiae (Sc) ... |
120-202 | 2.85e-06 | |||||||||||
Proteins containing this p23_like domain include p23 and its Saccharomyces cerevisiae (Sc) homolog Sba1. Both are co-chaperones for the heat shock protein (Hsp) 90. p23 binds Hsp90 and participates in the folding of a number of Hsp90 clients, including the progesterone receptor. p23 also has a passive chaperoning activity and in addition may participate in prostaglandin synthesis. Both p23 and Sba1p can regulate telomerase activity. This group includes domains similar to the C-terminal CHORD-SGT1 (CS) domain of suppressor of G2 allele of Skp1 (Sgt1). Sgt1 interacts with multiple protein complexes and has the features of a co-chaperone. Human (h) Sgt1 interacts with both Hsp70 and Hsp90, and has been shown to bind Hsp90 through its CS domain. Saccharomyces cerevisiae (Sc) Sgt1 is a subunit of both core kinetochore and SCF (Skp1-Cul1-F-box) ubiquitin ligase complexes. Sgt1 is required for pathogen resistance in plants. This group also includes the p23_like domains of human butyrate-induced transcript 1 (hB-ind1), NUD (nuclear distribution) C, Melusin, and NAD(P)H cytochrome b5 (NCB5) oxidoreductase (OR). hB-ind1 plays a role in the signaling pathway mediated by the small GTPase Rac1, NUDC is needed for nuclear movement, Melusin interacts with two splice variants of beta1 integrin, and NCB5OR plays a part in maintaining viable pancreatic beta cells. Pssm-ID: 107220 Cd Length: 84 Bit Score: 46.51 E-value: 2.85e-06
|
|||||||||||||||
| CS | pfam04969 | CS domain; The CS and CHORD (pfam04968) are fused into a single polypeptide chain in metazoans ... |
387-476 | 5.98e-06 | |||||||||||
CS domain; The CS and CHORD (pfam04968) are fused into a single polypeptide chain in metazoans but are found in separate proteins in plants; this is thought to be indicative of an interaction between CS and CHORD. It has been suggested that the CS domain is a binding module for HSP90, implying that CS domain-containing proteins are involved in recruiting heat shock proteins to multiprotein assemblies. Two CS domains are found at the N-terminus of Ubiquitin carboxyl-terminal hydrolase 19 (USP19), these domains may play a role in the interaction of USP19 with cellular inhibitor of apoptosis 2. Pssm-ID: 461503 Cd Length: 76 Bit Score: 45.33 E-value: 5.98e-06
|
|||||||||||||||
| PLN03088 | PLN03088 | SGT1, suppressor of G2 allele of SKP1; Provisional |
356-536 | 9.89e-05 | |||||||||||
SGT1, suppressor of G2 allele of SKP1; Provisional Pssm-ID: 215568 [Multi-domain] Cd Length: 356 Bit Score: 46.32 E-value: 9.89e-05
|
|||||||||||||||
| CS | pfam04969 | CS domain; The CS and CHORD (pfam04968) are fused into a single polypeptide chain in metazoans ... |
119-191 | 7.62e-03 | |||||||||||
CS domain; The CS and CHORD (pfam04968) are fused into a single polypeptide chain in metazoans but are found in separate proteins in plants; this is thought to be indicative of an interaction between CS and CHORD. It has been suggested that the CS domain is a binding module for HSP90, implying that CS domain-containing proteins are involved in recruiting heat shock proteins to multiprotein assemblies. Two CS domains are found at the N-terminus of Ubiquitin carboxyl-terminal hydrolase 19 (USP19), these domains may play a role in the interaction of USP19 with cellular inhibitor of apoptosis 2. Pssm-ID: 461503 Cd Length: 76 Bit Score: 36.85 E-value: 7.62e-03
|
|||||||||||||||
| Name | Accession | Description | Interval | E-value | |||||||||||
| UBP12 | COG5560 | Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; |
597-1319 | 1.21e-118 | |||||||||||
Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; Pssm-ID: 227847 [Multi-domain] Cd Length: 823 Bit Score: 391.55 E-value: 1.21e-118
|
|||||||||||||||
| USP19_linker | pfam16602 | Linker region of USP19 deubiquitinase; This region is generally located between a CS domain ... |
477-599 | 5.74e-58 | |||||||||||
Linker region of USP19 deubiquitinase; This region is generally located between a CS domain pfam04969 and the enzymatic UCH domain pfam00582 of USP19 deubiquitinases. This region is predicted to be natively unstructured. Its precise functional role is not known. Pssm-ID: 465191 Cd Length: 121 Bit Score: 195.52 E-value: 5.74e-58
|
|||||||||||||||
| Peptidase_C19R | cd02674 | A subfamily of peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1159-1316 | 6.15e-56 | |||||||||||
A subfamily of peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239139 [Multi-domain] Cd Length: 230 Bit Score: 194.04 E-value: 6.15e-56
|
|||||||||||||||
| UCH | pfam00443 | Ubiquitin carboxyl-terminal hydrolase; |
601-788 | 3.19e-48 | |||||||||||
Ubiquitin carboxyl-terminal hydrolase; Pssm-ID: 425685 [Multi-domain] Cd Length: 310 Bit Score: 174.94 E-value: 3.19e-48
|
|||||||||||||||
| UCH | pfam00443 | Ubiquitin carboxyl-terminal hydrolase; |
1164-1315 | 1.52e-46 | |||||||||||
Ubiquitin carboxyl-terminal hydrolase; Pssm-ID: 425685 [Multi-domain] Cd Length: 310 Bit Score: 169.93 E-value: 1.52e-46
|
|||||||||||||||
| Peptidase_C19 | cd02257 | Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ... |
1163-1316 | 1.88e-40 | |||||||||||
Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyse bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239072 [Multi-domain] Cd Length: 255 Bit Score: 150.33 E-value: 1.88e-40
|
|||||||||||||||
| Peptidase_C19D | cd02660 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1151-1315 | 2.68e-35 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239125 [Multi-domain] Cd Length: 328 Bit Score: 137.89 E-value: 2.68e-35
|
|||||||||||||||
| Peptidase_C19E | cd02661 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1163-1315 | 3.09e-33 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239126 [Multi-domain] Cd Length: 304 Bit Score: 131.24 E-value: 3.09e-33
|
|||||||||||||||
| Peptidase_C19K | cd02667 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1162-1316 | 2.44e-29 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239132 [Multi-domain] Cd Length: 279 Bit Score: 119.03 E-value: 2.44e-29
|
|||||||||||||||
| Peptidase_C19R | cd02674 | A subfamily of peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
602-788 | 4.99e-26 | |||||||||||
A subfamily of peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239139 [Multi-domain] Cd Length: 230 Bit Score: 108.14 E-value: 4.99e-26
|
|||||||||||||||
| peptidase_C19C | cd02659 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1168-1319 | 9.04e-25 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239124 [Multi-domain] Cd Length: 334 Bit Score: 107.34 E-value: 9.04e-25
|
|||||||||||||||
| Peptidase_C19G | cd02663 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1161-1315 | 6.26e-22 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239128 [Multi-domain] Cd Length: 300 Bit Score: 98.15 E-value: 6.26e-22
|
|||||||||||||||
| Peptidase_C19 | cd02257 | Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ... |
602-785 | 2.54e-21 | |||||||||||
Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyse bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239072 [Multi-domain] Cd Length: 255 Bit Score: 95.24 E-value: 2.54e-21
|
|||||||||||||||
| p23_CS_SGT1_like | cd06466 | p23_like domain similar to the C-terminal CHORD-SGT1 (CS) domain of Sgt1 (suppressor of G2 ... |
390-486 | 4.69e-21 | |||||||||||
p23_like domain similar to the C-terminal CHORD-SGT1 (CS) domain of Sgt1 (suppressor of G2 allele of Skp1). Sgt1 interacts with multiple protein complexes and has the features of a cochaperone. Human (h) Sgt1 interacts with both Hsp70 and Hsp90, and has been shown to bind Hsp90 through its CS domain. Saccharomyces cerevisiae (Sc) Sgt1 is a subunit of both core kinetochore and SCF (Skp1-Cul1-F-box) ubiquitin ligase complexes. Sgt1 is required for pathogen resistance in plants. ScSgt1 is needed for the G1/S and G2/M cell-cycle transitions, and for assembly of the core kinetochore complex (CBF3) via activation of Ctf13, the F-box protein. Binding of Hsp82 (a yeast Hsp90 homologue) to ScSgt1, promotes the binding of Sgt1 to Skp1 and of Skp1 to Ctf13. Some proteins in this group have an SGT1-specific (SGS) domain at the extreme C-terminus. The ScSgt1-SGS domain binds adenylate cyclase. The hSgt1-SGS domain interacts with some S100 family proteins, and studies suggest that the interaction of hSgt1 with Hsp90 and Hsp70 may be regulated by S100A6 in a Ca2+ dependent fashion. This group also includes the p23_like domains of Melusin and NAD(P)H cytochrome b5 (NCB5) oxidoreductase (OR). Melusin is a vertebrate protein which interacts with two splice variants of beta1 integrin, and NCB5OR plays a part in maintaining viable pancreatic beta cells. Pssm-ID: 107223 Cd Length: 84 Bit Score: 88.80 E-value: 4.69e-21
|
|||||||||||||||
| Peptidase_C19E | cd02661 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
601-778 | 7.01e-21 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239126 [Multi-domain] Cd Length: 304 Bit Score: 95.04 E-value: 7.01e-21
|
|||||||||||||||
| Peptidase_C19D | cd02660 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
602-783 | 4.17e-20 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239125 [Multi-domain] Cd Length: 328 Bit Score: 93.21 E-value: 4.17e-20
|
|||||||||||||||
| Peptidase_C19L | cd02668 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1159-1316 | 3.22e-19 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239133 [Multi-domain] Cd Length: 324 Bit Score: 90.56 E-value: 3.22e-19
|
|||||||||||||||
| Peptidase_C19H | cd02664 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1168-1316 | 1.83e-16 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239129 [Multi-domain] Cd Length: 327 Bit Score: 82.15 E-value: 1.83e-16
|
|||||||||||||||
| Peptidase_C19K | cd02667 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
602-787 | 3.01e-16 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239132 [Multi-domain] Cd Length: 279 Bit Score: 80.89 E-value: 3.01e-16
|
|||||||||||||||
| Peptidase_C19H | cd02664 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
602-795 | 1.48e-15 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239129 [Multi-domain] Cd Length: 327 Bit Score: 79.46 E-value: 1.48e-15
|
|||||||||||||||
| Peptidase_C19O | cd02671 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1131-1316 | 4.90e-15 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239136 [Multi-domain] Cd Length: 332 Bit Score: 78.01 E-value: 4.90e-15
|
|||||||||||||||
| Peptidase_C19M | cd02669 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
597-783 | 4.16e-14 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239134 [Multi-domain] Cd Length: 440 Bit Score: 76.59 E-value: 4.16e-14
|
|||||||||||||||
| Peptidase_C19A | cd02657 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1191-1316 | 3.46e-12 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyse bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239122 [Multi-domain] Cd Length: 305 Bit Score: 68.90 E-value: 3.46e-12
|
|||||||||||||||
| zf-MYND | pfam01753 | MYND finger; |
895-937 | 6.37e-12 | |||||||||||
MYND finger; Pssm-ID: 460312 Cd Length: 39 Bit Score: 61.28 E-value: 6.37e-12
|
|||||||||||||||
| COG5533 | COG5533 | Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; |
602-726 | 8.11e-12 | |||||||||||
Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; Pssm-ID: 444284 [Multi-domain] Cd Length: 284 Bit Score: 67.52 E-value: 8.11e-12
|
|||||||||||||||
| COG5533 | COG5533 | Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; |
1163-1317 | 1.19e-11 | |||||||||||
Ubiquitin C-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; Pssm-ID: 444284 [Multi-domain] Cd Length: 284 Bit Score: 67.13 E-value: 1.19e-11
|
|||||||||||||||
| COG5077 | COG5077 | Ubiquitin carboxyl-terminal hydrolase [Posttranslational modification, protein turnover, ... |
1163-1305 | 1.38e-11 | |||||||||||
Ubiquitin carboxyl-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; Pssm-ID: 227409 [Multi-domain] Cd Length: 1089 Bit Score: 69.51 E-value: 1.38e-11
|
|||||||||||||||
| Peptidase_C19G | cd02663 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
602-782 | 3.12e-11 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239128 [Multi-domain] Cd Length: 300 Bit Score: 66.18 E-value: 3.12e-11
|
|||||||||||||||
| Peptidase_C19F | cd02662 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1163-1315 | 5.12e-11 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239127 [Multi-domain] Cd Length: 240 Bit Score: 64.31 E-value: 5.12e-11
|
|||||||||||||||
| Peptidase_C19A | cd02657 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
602-760 | 8.06e-11 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyse bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239122 [Multi-domain] Cd Length: 305 Bit Score: 65.04 E-value: 8.06e-11
|
|||||||||||||||
| Peptidase_C19B | cd02658 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
602-780 | 5.03e-10 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239123 [Multi-domain] Cd Length: 311 Bit Score: 62.34 E-value: 5.03e-10
|
|||||||||||||||
| Peptidase_C19B | cd02658 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1163-1316 | 1.24e-09 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239123 [Multi-domain] Cd Length: 311 Bit Score: 61.18 E-value: 1.24e-09
|
|||||||||||||||
| p23_like | cd06463 | Proteins containing this p23_like domain include p23 and its Saccharomyces cerevisiae (Sc) ... |
396-486 | 2.74e-09 | |||||||||||
Proteins containing this p23_like domain include p23 and its Saccharomyces cerevisiae (Sc) homolog Sba1. Both are co-chaperones for the heat shock protein (Hsp) 90. p23 binds Hsp90 and participates in the folding of a number of Hsp90 clients, including the progesterone receptor. p23 also has a passive chaperoning activity and in addition may participate in prostaglandin synthesis. Both p23 and Sba1p can regulate telomerase activity. This group includes domains similar to the C-terminal CHORD-SGT1 (CS) domain of suppressor of G2 allele of Skp1 (Sgt1). Sgt1 interacts with multiple protein complexes and has the features of a co-chaperone. Human (h) Sgt1 interacts with both Hsp70 and Hsp90, and has been shown to bind Hsp90 through its CS domain. Saccharomyces cerevisiae (Sc) Sgt1 is a subunit of both core kinetochore and SCF (Skp1-Cul1-F-box) ubiquitin ligase complexes. Sgt1 is required for pathogen resistance in plants. This group also includes the p23_like domains of human butyrate-induced transcript 1 (hB-ind1), NUD (nuclear distribution) C, Melusin, and NAD(P)H cytochrome b5 (NCB5) oxidoreductase (OR). hB-ind1 plays a role in the signaling pathway mediated by the small GTPase Rac1, NUDC is needed for nuclear movement, Melusin interacts with two splice variants of beta1 integrin, and NCB5OR plays a part in maintaining viable pancreatic beta cells. Pssm-ID: 107220 Cd Length: 84 Bit Score: 55.37 E-value: 2.74e-09
|
|||||||||||||||
| Peptidase_C19O | cd02671 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
573-706 | 2.30e-07 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239136 [Multi-domain] Cd Length: 332 Bit Score: 54.51 E-value: 2.30e-07
|
|||||||||||||||
| peptidase_C19C | cd02659 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
599-778 | 3.00e-07 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239124 [Multi-domain] Cd Length: 334 Bit Score: 54.19 E-value: 3.00e-07
|
|||||||||||||||
| Peptidase_C19F | cd02662 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
602-628 | 9.88e-07 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239127 [Multi-domain] Cd Length: 240 Bit Score: 51.60 E-value: 9.88e-07
|
|||||||||||||||
| p23_like | cd06463 | Proteins containing this p23_like domain include p23 and its Saccharomyces cerevisiae (Sc) ... |
120-202 | 2.85e-06 | |||||||||||
Proteins containing this p23_like domain include p23 and its Saccharomyces cerevisiae (Sc) homolog Sba1. Both are co-chaperones for the heat shock protein (Hsp) 90. p23 binds Hsp90 and participates in the folding of a number of Hsp90 clients, including the progesterone receptor. p23 also has a passive chaperoning activity and in addition may participate in prostaglandin synthesis. Both p23 and Sba1p can regulate telomerase activity. This group includes domains similar to the C-terminal CHORD-SGT1 (CS) domain of suppressor of G2 allele of Skp1 (Sgt1). Sgt1 interacts with multiple protein complexes and has the features of a co-chaperone. Human (h) Sgt1 interacts with both Hsp70 and Hsp90, and has been shown to bind Hsp90 through its CS domain. Saccharomyces cerevisiae (Sc) Sgt1 is a subunit of both core kinetochore and SCF (Skp1-Cul1-F-box) ubiquitin ligase complexes. Sgt1 is required for pathogen resistance in plants. This group also includes the p23_like domains of human butyrate-induced transcript 1 (hB-ind1), NUD (nuclear distribution) C, Melusin, and NAD(P)H cytochrome b5 (NCB5) oxidoreductase (OR). hB-ind1 plays a role in the signaling pathway mediated by the small GTPase Rac1, NUDC is needed for nuclear movement, Melusin interacts with two splice variants of beta1 integrin, and NCB5OR plays a part in maintaining viable pancreatic beta cells. Pssm-ID: 107220 Cd Length: 84 Bit Score: 46.51 E-value: 2.85e-06
|
|||||||||||||||
| CS | pfam04969 | CS domain; The CS and CHORD (pfam04968) are fused into a single polypeptide chain in metazoans ... |
387-476 | 5.98e-06 | |||||||||||
CS domain; The CS and CHORD (pfam04968) are fused into a single polypeptide chain in metazoans but are found in separate proteins in plants; this is thought to be indicative of an interaction between CS and CHORD. It has been suggested that the CS domain is a binding module for HSP90, implying that CS domain-containing proteins are involved in recruiting heat shock proteins to multiprotein assemblies. Two CS domains are found at the N-terminus of Ubiquitin carboxyl-terminal hydrolase 19 (USP19), these domains may play a role in the interaction of USP19 with cellular inhibitor of apoptosis 2. Pssm-ID: 461503 Cd Length: 76 Bit Score: 45.33 E-value: 5.98e-06
|
|||||||||||||||
| Peptidase_C19I | cd02665 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1203-1315 | 1.89e-05 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239130 [Multi-domain] Cd Length: 228 Bit Score: 47.55 E-value: 1.89e-05
|
|||||||||||||||
| Peptidase_C19Q | cd02673 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1185-1315 | 6.38e-05 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239138 [Multi-domain] Cd Length: 245 Bit Score: 46.37 E-value: 6.38e-05
|
|||||||||||||||
| PLN03088 | PLN03088 | SGT1, suppressor of G2 allele of SKP1; Provisional |
356-536 | 9.89e-05 | |||||||||||
SGT1, suppressor of G2 allele of SKP1; Provisional Pssm-ID: 215568 [Multi-domain] Cd Length: 356 Bit Score: 46.32 E-value: 9.89e-05
|
|||||||||||||||
| Peptidase_C19M | cd02669 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1193-1264 | 1.18e-04 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239134 [Multi-domain] Cd Length: 440 Bit Score: 46.16 E-value: 1.18e-04
|
|||||||||||||||
| Peptidase_C19J | cd02666 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
601-801 | 1.29e-04 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239131 [Multi-domain] Cd Length: 343 Bit Score: 45.95 E-value: 1.29e-04
|
|||||||||||||||
| COG5077 | COG5077 | Ubiquitin carboxyl-terminal hydrolase [Posttranslational modification, protein turnover, ... |
599-704 | 3.17e-04 | |||||||||||
Ubiquitin carboxyl-terminal hydrolase [Posttranslational modification, protein turnover, chaperones]; Pssm-ID: 227409 [Multi-domain] Cd Length: 1089 Bit Score: 45.25 E-value: 3.17e-04
|
|||||||||||||||
| p23_CS_hSgt1_like | cd06489 | p23_like domain similar to the C-terminal CS (CHORD-SGT1) domain of human (h) Sgt1 and related ... |
399-486 | 3.43e-04 | |||||||||||
p23_like domain similar to the C-terminal CS (CHORD-SGT1) domain of human (h) Sgt1 and related proteins. hSgt1 is a co-chaperone which has been shown to be elevated in HEp-2 cells as a result of stress conditions such as heat shock. It interacts with the heat shock proteins (HSPs) Hsp70 and Hsp90, and it expression pattern is synchronized with these two Hsps. The interaction with HSP90 has been shown to involve the hSgt1_CS domain, and appears to be required for correct kinetochore assembly and efficient cell division. Some proteins in this subgroup contain a tetratricopeptide repeat (TPR) HSP-binding domain N-terminal to this CS domain, and most proteins in this subgroup contain a Sgt1-specific (SGS) domain C-terminal to the CS domain. The SGS domain interacts with some S100 family proteins. Studies suggest that S100A6 modulates in a Ca2+ dependent manner the interactions of hSgt1 with Hsp90 and Hsp70. The yeast Sgt1 CS domain is not found in this subgroup. Pssm-ID: 107239 Cd Length: 84 Bit Score: 40.82 E-value: 3.43e-04
|
|||||||||||||||
| USP7_C2 | pfam14533 | Ubiquitin-specific protease C-terminal; This C-terminal domain on many long ubiquitin-specific ... |
772-861 | 4.03e-04 | |||||||||||
Ubiquitin-specific protease C-terminal; This C-terminal domain on many long ubiquitin-specific proteases has no known function. Pssm-ID: 464201 Cd Length: 204 Bit Score: 43.24 E-value: 4.03e-04
|
|||||||||||||||
| p23_NUDC_like | cd06467 | p23_like domain of NUD (nuclear distribution) C and similar proteins. Aspergillus nidulas (An) ... |
120-202 | 4.96e-04 | |||||||||||
p23_like domain of NUD (nuclear distribution) C and similar proteins. Aspergillus nidulas (An) NUDC is needed for nuclear movement. AnNUDC is localized at the hyphal cortex, and binds NUDF at spindle pole bodies (SPBs) and in the cytoplasm at different stages in the cell cycle. At the SPBs it is part of the dynein molecular motor/NUDF complex that regulates microtubule dynamics. Mammalian(m) NUDC associates both with the dynein complex and also with an anti-inflammatory enzyme, platelet activating factor acetylhydrolase I, PAF-AH(I) complex, through binding mNUDF, the regulatory beta subunit of PAF-AH(I). mNUDC is important for cell proliferation both in normal and tumor tissues. Its expression is elevated in various cell types undergoing mitosis or stimulated to proliferate, with high expression levels observed in leukemic cells and tumors. For a leukemic cell line, human NUDC was shown to activate the thrombopoietin (TPO) receptor (Mpl) by binding to its extracellular domain, and promoting cell proliferation and differentiation. This group also includes the human broadly immunogenic tumor associated antigen, CML66, which is highly expressed in a variety of solid tumors and in leukemias. In normal tissues high expression of CML66 is limited to testis and heart. Pssm-ID: 107224 Cd Length: 85 Bit Score: 40.22 E-value: 4.96e-04
|
|||||||||||||||
| p23_melusin_like | cd06488 | p23_like domain similar to the C-terminal (tail) domain of vertebrate Melusin and related ... |
398-486 | 5.50e-04 | |||||||||||
p23_like domain similar to the C-terminal (tail) domain of vertebrate Melusin and related proteins. Melusin's tail domain interacts with the cytoplasmic domain of beta1-A and beta1-D isoforms of beta1 integrin, it does not bind other integrin beta subunits. Melusin is a muscle-specific protein expressed in skeletal and cardiac muscles but not in smooth muscle or other tissues. It is needed for heart hypertrophy following mechanical overload. The integrin-binding portion of this domain appears to be sequestered in the full length melusin protein, Ca2+ may modulate the protein's conformation exposing this binding site. This group includes Chordc1, also known as Chp-1, which is conserved from vertebrates to humans. Mammalian Chordc1 interacts with the heat shock protein (HSP) Hsp90 and is implicated in circadian and/or homeostatic mechanisms in the brain. The N-terminal portions of proteins belonging to this group contain two cysteine and histidine rich domain (CHORD) domains. Pssm-ID: 107238 Cd Length: 87 Bit Score: 40.36 E-value: 5.50e-04
|
|||||||||||||||
| Peptidase_C19P | cd02672 | A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are ... |
1166-1315 | 6.12e-03 | |||||||||||
A subfamily of Peptidase C19. Peptidase C19 contains ubiquitinyl hydrolases. They are intracellular peptidases that remove ubiquitin molecules from polyubiquinated peptides by cleavage of isopeptide bonds. They hydrolyze bonds involving the carboxyl group of the C-terminal Gly residue of ubiquitin. The purpose of the de-ubiquitination is thought to be editing of the ubiquitin conjugates, which could rescue them from degradation, as well as recycling of the ubiquitin. The ubiquitin/proteasome system is responsible for most protein turnover in the mammalian cell, and with over 50 members, family C19 is one of the largest families of peptidases in the human genome. Pssm-ID: 239137 [Multi-domain] Cd Length: 268 Bit Score: 40.19 E-value: 6.12e-03
|
|||||||||||||||
| CS | pfam04969 | CS domain; The CS and CHORD (pfam04968) are fused into a single polypeptide chain in metazoans ... |
119-191 | 7.62e-03 | |||||||||||
CS domain; The CS and CHORD (pfam04968) are fused into a single polypeptide chain in metazoans but are found in separate proteins in plants; this is thought to be indicative of an interaction between CS and CHORD. It has been suggested that the CS domain is a binding module for HSP90, implying that CS domain-containing proteins are involved in recruiting heat shock proteins to multiprotein assemblies. Two CS domains are found at the N-terminus of Ubiquitin carboxyl-terminal hydrolase 19 (USP19), these domains may play a role in the interaction of USP19 with cellular inhibitor of apoptosis 2. Pssm-ID: 461503 Cd Length: 76 Bit Score: 36.85 E-value: 7.62e-03
|
|||||||||||||||
| Blast search parameters | ||||
|